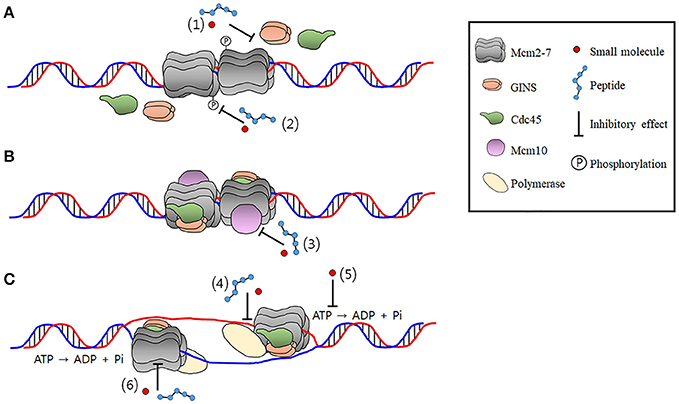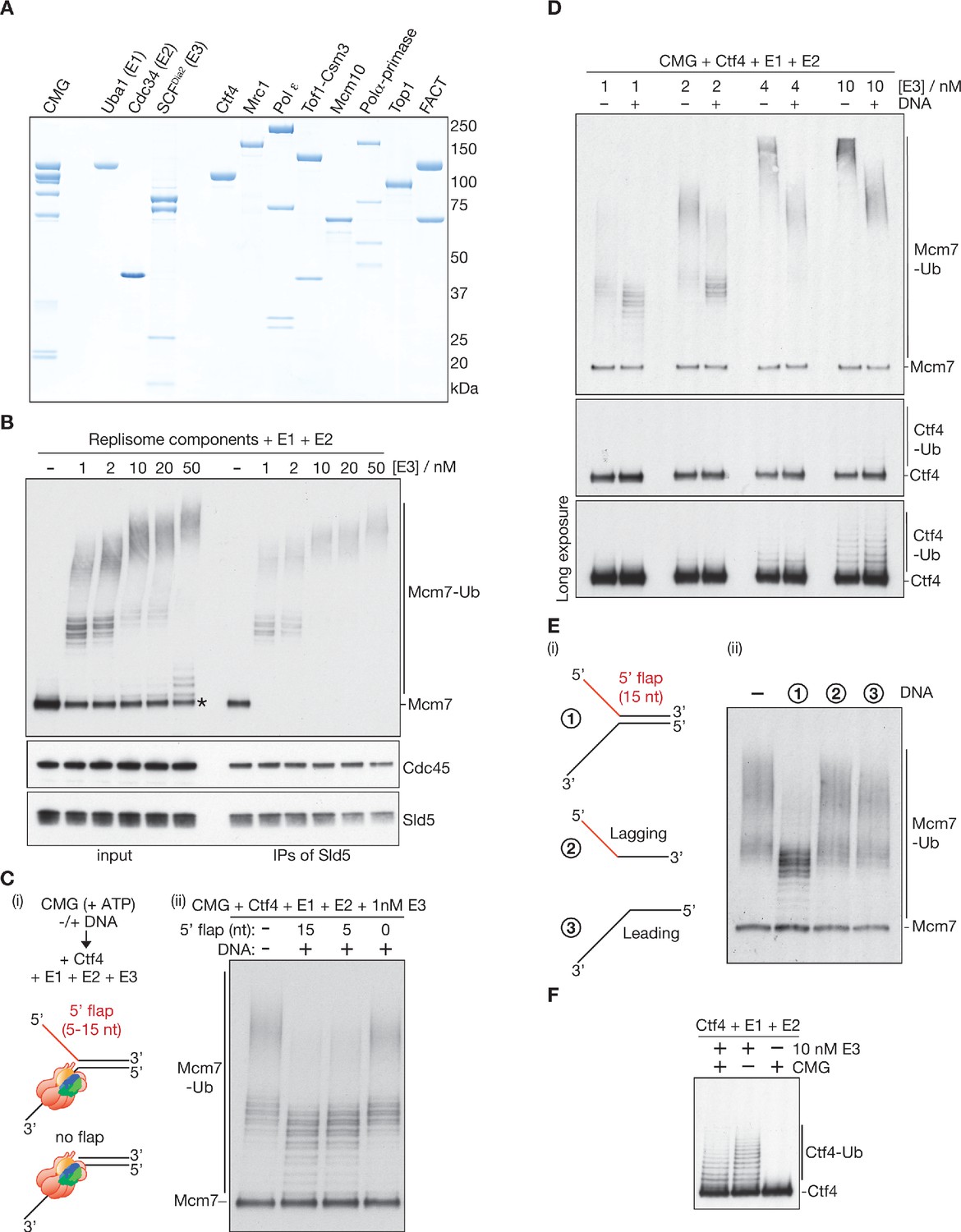
CMG helicase disassembly is controlled by replication fork DNA, replisome components and a ubiquitin threshold | eLife

Replication Fork Activation Is Enabled by a Single-Stranded DNA Gate in CMG Helicase - ScienceDirect

Mcm10 regulates the assembly of the CMG complex and DNA origin melting.... | Download Scientific Diagram

DNA densities and interactions with CMG in the CMG-ssDNA structure. CMG... | Download Scientific Diagram

Purification of the CMG complex. (A) Purification schematic. See the... | Download Scientific Diagram
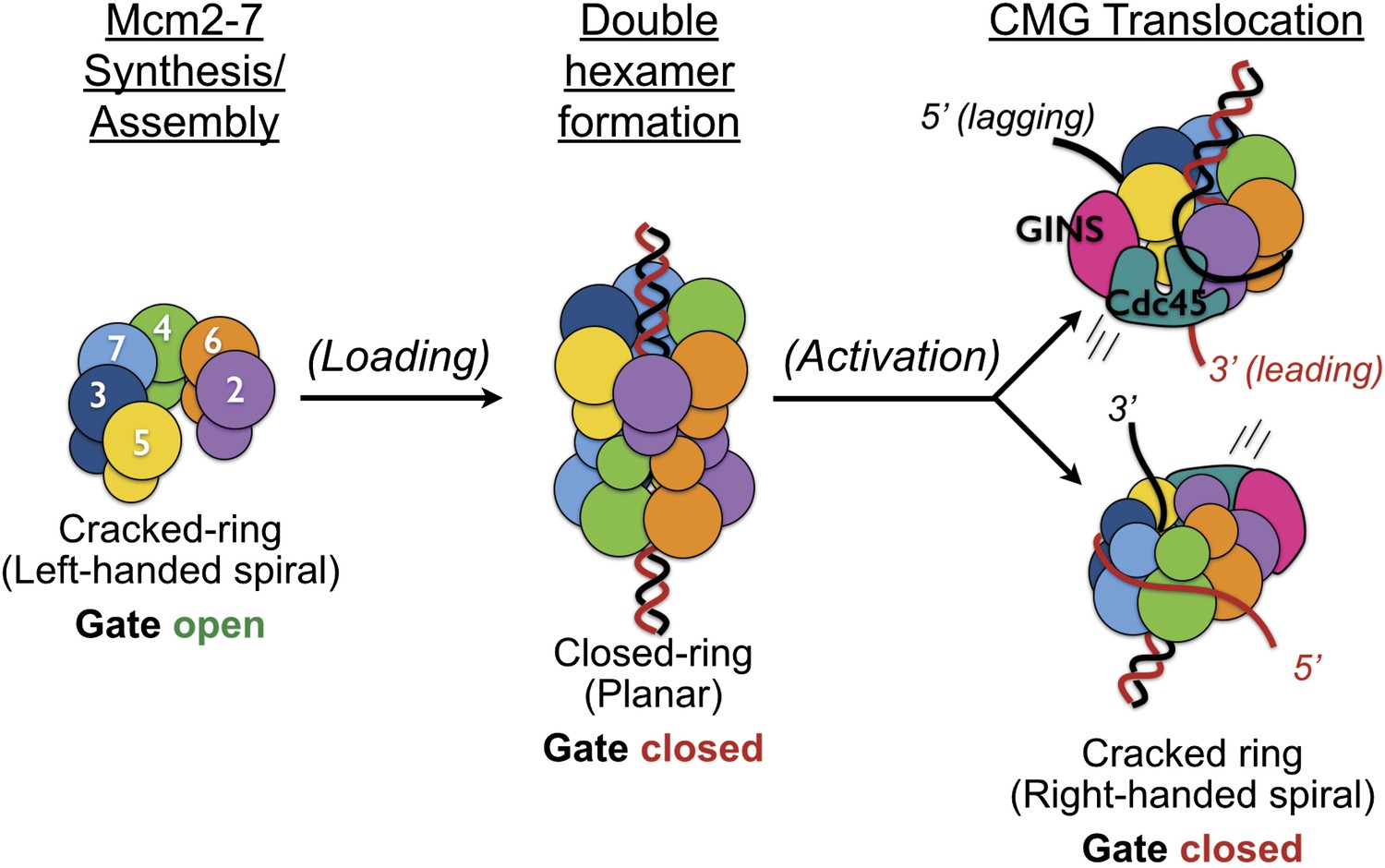
DNA binding polarity, dimerization, and ATPase ring remodeling in the CMG helicase of the eukaryotic replisome | eLife
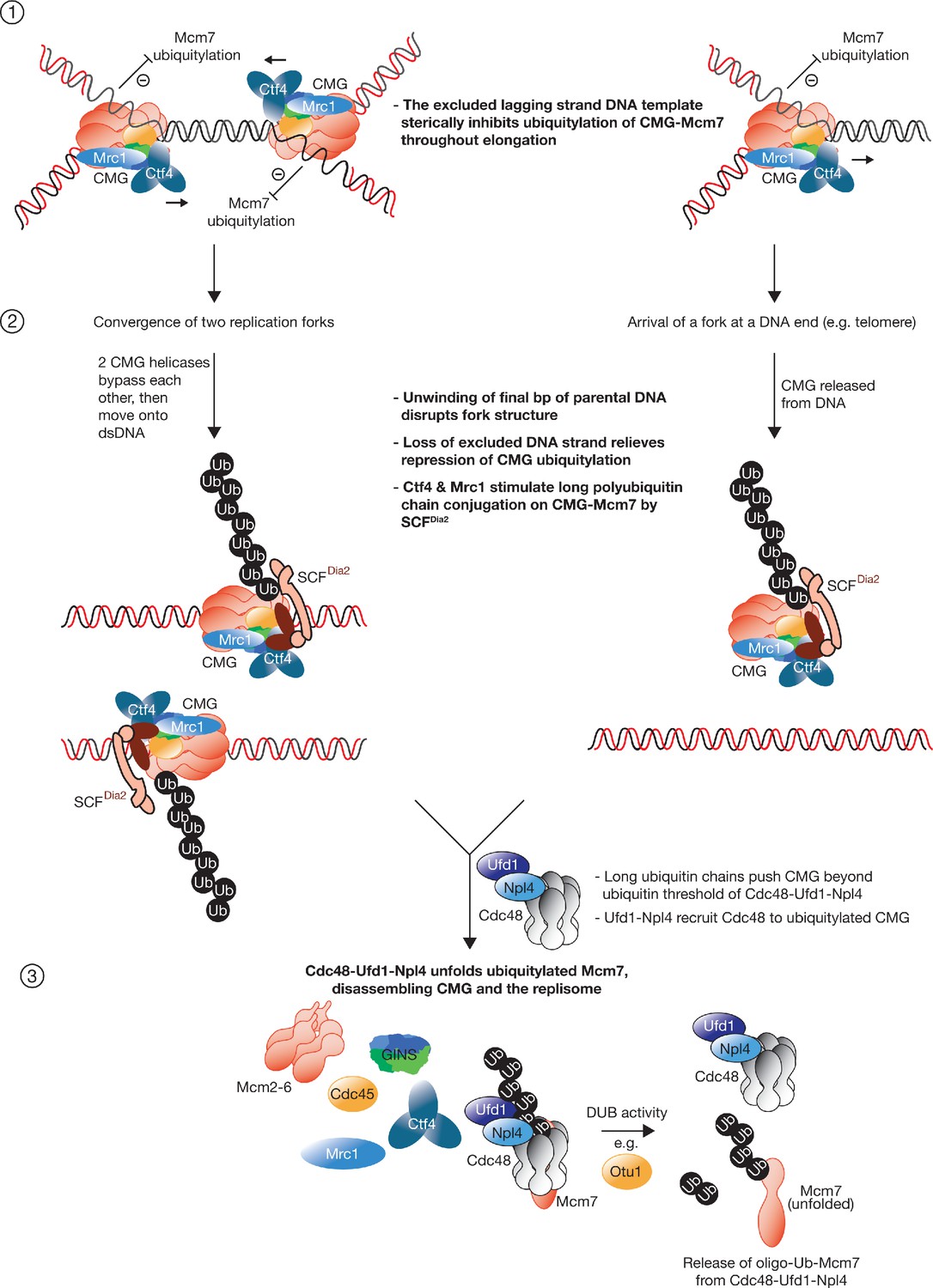
CMG helicase disassembly is controlled by replication fork DNA, replisome components and a ubiquitin threshold | eLife
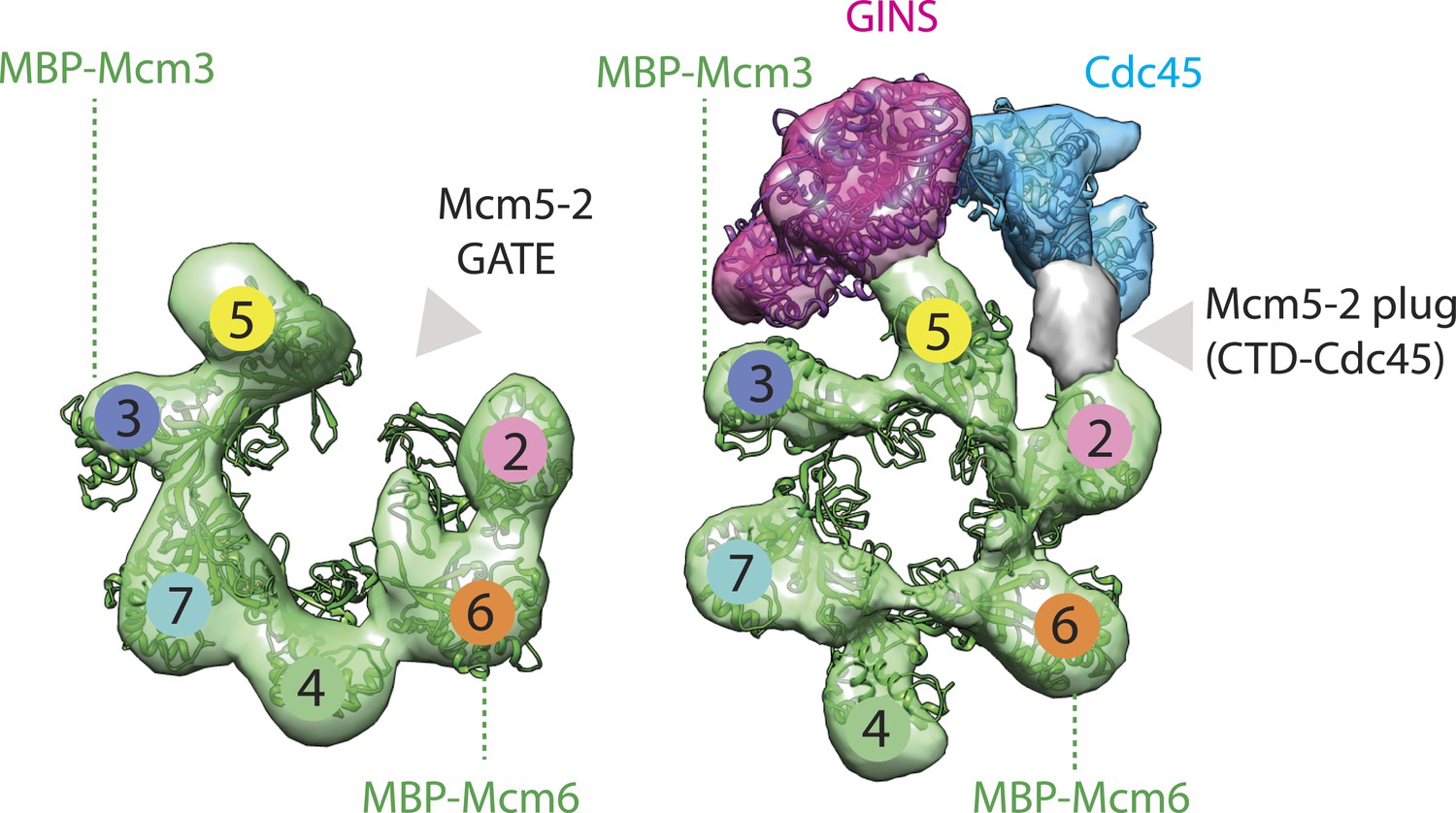
DNA binding polarity, dimerization, and ATPase ring remodeling in the CMG helicase of the eukaryotic replisome | eLife

Structure of eukaryotic CMG helicase at a replication fork and implications to replisome architecture and origin initiation | PNAS

Mechanisms in helicase activation and analysis of the replication fork activity and architecture – Speck Lab / DNA Replication Group – Christian Speck
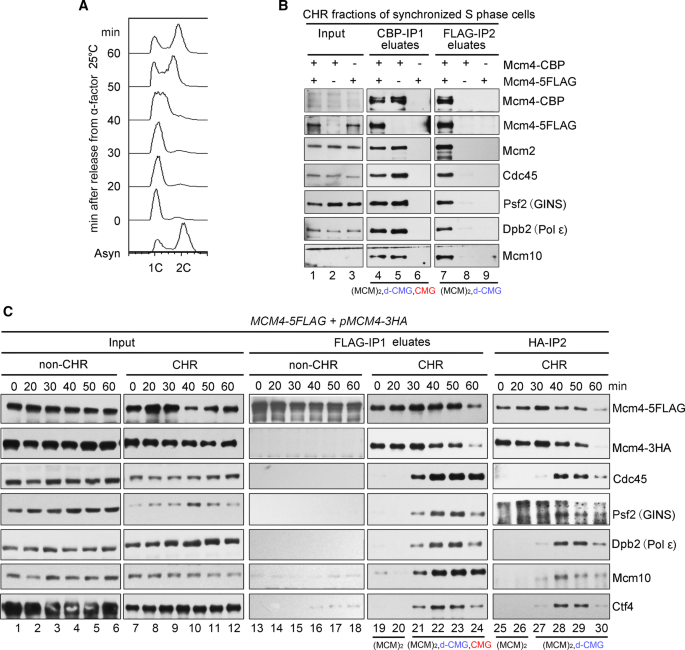
Characterization of the dimeric CMG/pre-initiation complex and its transition into DNA replication forks | SpringerLink

CMG helicase structure and translocation mechanism. (a) Low resolution... | Download Scientific Diagram
Structural components of the replisome. The CMG replicative helicase... | Download Scientific Diagram
Isolation of the Cdc45 Mcm2–7 GINS (CMG) complex, a candidate for the eukaryotic DNA replication fork helicase
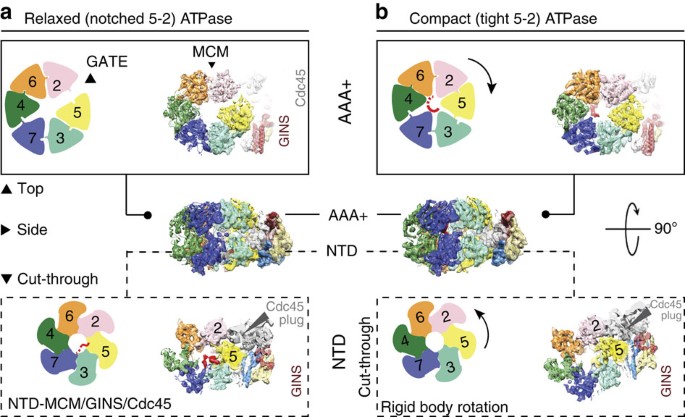
Cryo-EM structures of the eukaryotic replicative helicase bound to a translocation substrate | Nature Communications